Research interests
RNA plays critical and diverse roles in the cell. Messenger RNAs (mRNAs) that encode proteins have untranslated regions (UTRs), which direct the translation of these messages. To dictate when the transcript is translated and how much protein is made, these UTRs serve as landing pads for protein factors that fine-tune translation accordingly. We aim to discover and explore the strategies that RNA-binding proteins use to recognize and functionalize their target RNAs. We integrate biophysics, biochemistry, and structural biology to develop and test mechanistic hypotheses; these tools yield quantitative data and snapshots of molecules in action, allowing us a glimpse of how systems function at the molecular level.
We have established the RNA-binding protein La-related protein 1 (LARP1) as a model system for understanding several fundamental biological principles, including RNA binding, protein-protein interactions, translational regulation, and the functional consequences of post-translational and post-transcriptional modifications.
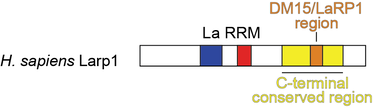
LARP1 has been implicated in several cancers, including that of the cervix, ovary, lung, and liver. Our collaborator, Dr. Sarah Blagden (University of Oxford) demonstrated that increased LARP1 expression correlates with disease progression in cervical cancer (Mura et al., 2014). Thus, understanding the structure, function, and binding partners of LARP1 will aid in designing diagnostic tools and new treatment targets, leading to effective, targeted therapies for various cancers.
LARP1 has been proposed to modulate the translation levels of specific proteins, perhaps through its association with poly(A) binding protein (PABP). PABP protects the 3’ poly(A) tail of mRNA from degradation and acts as a scaffold upon which other proteins assemble to regulate mRNA metabolism . The protein-protein interactions fostered by PABP boost translation initiation and ribosomal recruitment, or repress translation by inhibiting initiation or by promoting mRNA turnover.
LARP1 has been proposed to modulate the translation levels of specific proteins, perhaps through its association with poly(A) binding protein (PABP). PABP protects the 3’ poly(A) tail of mRNA from degradation and acts as a scaffold upon which other proteins assemble to regulate mRNA metabolism . The protein-protein interactions fostered by PABP boost translation initiation and ribosomal recruitment, or repress translation by inhibiting initiation or by promoting mRNA turnover.
While we do not understand all of its roles in the cell, we have provided critical mechanistic insights that begin to define its molecular roles.
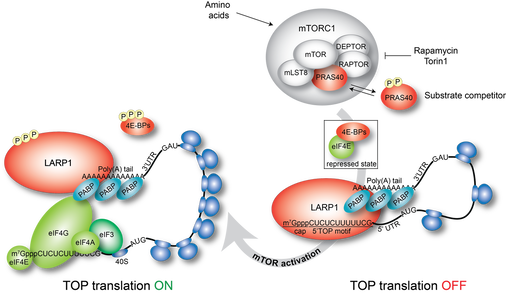
LARP1 has been identified as a node in the mTORC1 signaling pathway, which controls cell growth and proliferation by stimulating the biogenesis of ribosomes. mTORC1 activates the translation of a class of mRNAs that encode all of the ribosomal proteins, many translation factors, and other select groups of mRNAs that are characterized by a tract of pyrimidine nucleotides at the very 5’ end of the 5’UTR, termed the 5’terminal oligopyrimidine (5’TOP) motif.
We determined that the C-terminal conserved region of LARP1, the DM15 region,binds the pyrimidine-rich sequence motif characteristic of TOP mRNAs (Lahr et al., 2015) and directly engages the 7-methylguanosine cap of bound TOP transcripts (Lahr et al., 2017); these activities of LARP1 preventing the assembly of the translation initiation factors on these transcripts. This study defined LARP1 as a bona-fide cap-binding protein that is likely a direct translational repressor of TOP transcripts—and therefore of ribosome biogenesis (reviewed in Fonseca et al., 2018).
Work in collaboration with Dr. Jacob Durrant's lab revealed the dynamics underlying capped-TOP recognition by the DM15 region of LARP1 (Cassidy, Lahr, et al., 2019). An exciting study from our lab also demonstrated that the La-Module of LARP1 has the ability to bind the 5'TOP motif and polyA RNA simultaneously (Al-Ashtal et al., 2019).
Future work on LARP1 will not only fill the gaps in our knowledge on its RNA- and protein- binding activities, but it will also reveal novel roles that RNA-binding proteins can play in cellular metabolism. Some questions of interest include:
Why does LARP1 bind some TOPs and not others?
How do post-translational modifications of the protein redefine these interactions or elicit new ones?
How does the interaction of LARP1 with PABP affect bound transcripts?
LARP1 has been identified as a node in the mTORC1 signaling pathway, which controls cell growth and proliferation by stimulating the biogenesis of ribosomes. mTORC1 activates the translation of a class of mRNAs that encode all of the ribosomal proteins, many translation factors, and other select groups of mRNAs that are characterized by a tract of pyrimidine nucleotides at the very 5’ end of the 5’UTR, termed the 5’terminal oligopyrimidine (5’TOP) motif.
We determined that the C-terminal conserved region of LARP1, the DM15 region,binds the pyrimidine-rich sequence motif characteristic of TOP mRNAs (Lahr et al., 2015) and directly engages the 7-methylguanosine cap of bound TOP transcripts (Lahr et al., 2017); these activities of LARP1 preventing the assembly of the translation initiation factors on these transcripts. This study defined LARP1 as a bona-fide cap-binding protein that is likely a direct translational repressor of TOP transcripts—and therefore of ribosome biogenesis (reviewed in Fonseca et al., 2018).
Work in collaboration with Dr. Jacob Durrant's lab revealed the dynamics underlying capped-TOP recognition by the DM15 region of LARP1 (Cassidy, Lahr, et al., 2019). An exciting study from our lab also demonstrated that the La-Module of LARP1 has the ability to bind the 5'TOP motif and polyA RNA simultaneously (Al-Ashtal et al., 2019).
Future work on LARP1 will not only fill the gaps in our knowledge on its RNA- and protein- binding activities, but it will also reveal novel roles that RNA-binding proteins can play in cellular metabolism. Some questions of interest include:
Why does LARP1 bind some TOPs and not others?
How do post-translational modifications of the protein redefine these interactions or elicit new ones?
How does the interaction of LARP1 with PABP affect bound transcripts?